PDF) A fast and interpretable deep learning approach for accurate electrostatics-driven pKa predictions in proteins
4.5 (528) · € 31.00 · In Magazzino


A Fast and Interpretable Deep Learning Approach for Accurate Electrostatics-Driven pKa Predictions in Proteins

Protein pKa Prediction with Machine Learning
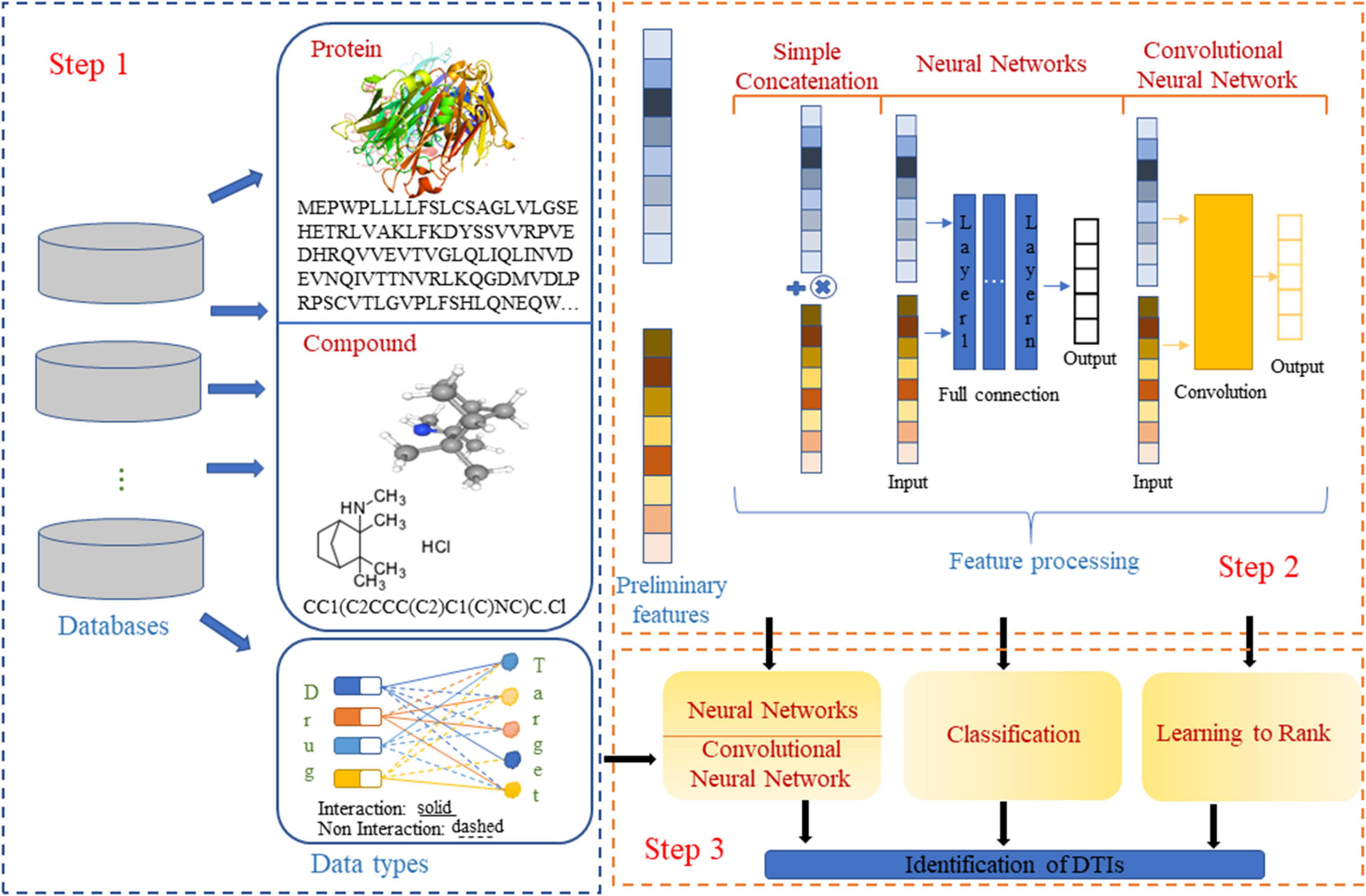
Frontiers, Application of Machine Learning for Drug–Target Interaction Prediction

Frontiers Improving Small Molecule pKa Prediction Using Transfer Learning With Graph Neural Networks
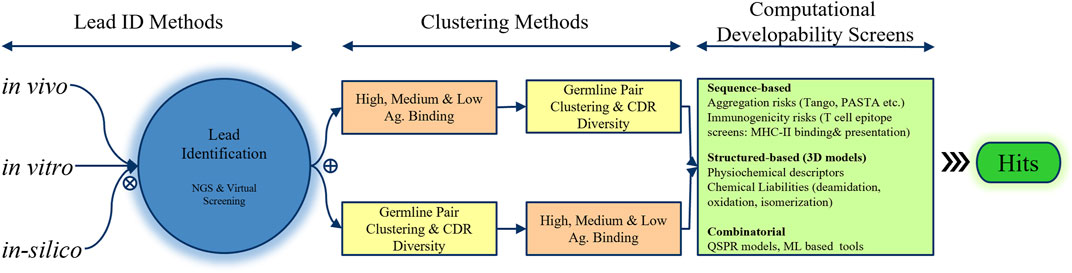
Frontiers How can we discover developable antibody-based biotherapeutics?

Comparative Performance of High-Throughput Methods for Protein pKa Predictions

Biomolecules, Free Full-Text

PDF) Protein pKa prediction by tree-based machine learning
![PDF] PKAD: a database of experimentally measured pKa values of ionizable groups in proteins](https://d3i71xaburhd42.cloudfront.net/fa39ae2bb4f763f948fe5edea27efbc9e4fa34d5/5-Figure2-1.png)
PDF] PKAD: a database of experimentally measured pKa values of ionizable groups in proteins

PDF) A Fast and Interpretable Deep Learning Approach for Accurate Electrostatics-Driven p K a Predictions in Proteins

Djork-Arné Clevert - Pfizer

Surface-based protein domains retrieval methods from a SHREC2021 challenge - ScienceDirect

Biophysical cartography of the native and human-engineered antibody landscapes quantifies the plasticity of antibody developability






![6 Tips to Achieve Flow State at Work [2023] • Asana](https://assets.asana.biz/transform/ec5b003f-4c31-4aab-80f2-96059caebfe4/article-productivity-flow-state-work-2x)
:max_bytes(150000):strip_icc()/021323-italian-bob-lead-f6cb3798704344de96199871507c7a05.jpg)